Non-coding RNAs and Brain Evolution
During their life-span neurons undergo profound changes in gene expression that trigger rapid transitions in their developmental trajectory or functional state. The overarching goal of the research in the Lippi lab is to identify the key molecular players that drive and coordinate these transcriptional changes. In particular, we focus on non-coding RNAs (ncRNAs), a novel and exciting class of master regulators of gene expression that emerged during evolution to confer robustness to biological systems. Intriguingly, recent discoveries have suggested that the protein-coding repertoire has not changed much across multicellular organisms, and that increases in biological complexity mostly derives from the expansion of the ncRNA repertoire (Figure 1)
Figure 1. Non-coding RNAs drive increases in complexity. Relationship between biological complexity (increasing number of different cell types) and genetic information (protein-coding or non-coding genes). The number of protein-coding genes for each organism (orange squares) has plateaued at the dawn of multicellularity: there is little or no increase from Dictyostelium to H. Sapiens. The number of non-coding genes (blue rhombi), instead, shows a remarkable linear relationship with the number of different cell types. This suggests that to achieve such a massive increase in biological complexity, evolution did not add more building blocks (proteins), but layers upon layers of sophisticated regulation of gene expression. Adapted from Liu et al. 2013.
Many ncRNAs are enriched in the brain and increase their expression during development, suggesting that they play fundamental roles in establishing properly balanced neural networks. Consistently, a wealth of recent literature indicates that changes in ncRNA levels are linked to multiple neurodevelopmental disorders, including autism, schizophrenia, and epilepsy.
microRNAs Instruct Neural Network Formation
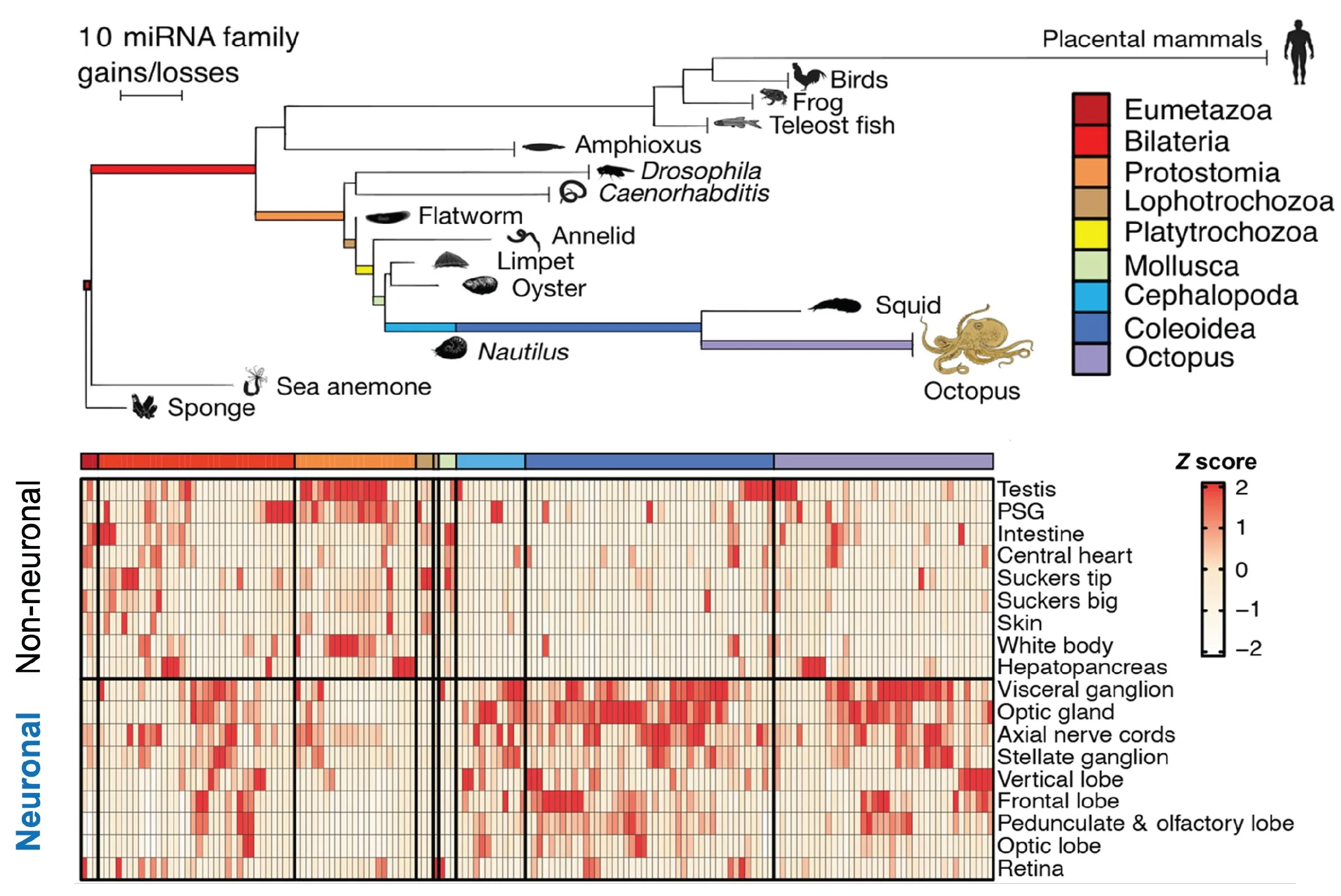
One of the most studied classes of ncRNAs is microRNAs, post-transcriptional regulators of gene expression that bind and repress mRNAs based on sequence complementarity to a microRNA response element (MRE). The microRNA repertoire strongly correlates with brain complexity, suggesting that increasing microRNA-mediated regulation of gene expression was critical for building and maintaining larger brains (Figure 2).
Figure 2. Emergence of new microRNA families. Rapid expansion of the microRNA repertoire in animals with large brains, such as primates and octopi (top). The majority of new microRNA families emerged in the octopus’ neuronal tissue (bottom). Adapted from Zolatorov et al. 2022.
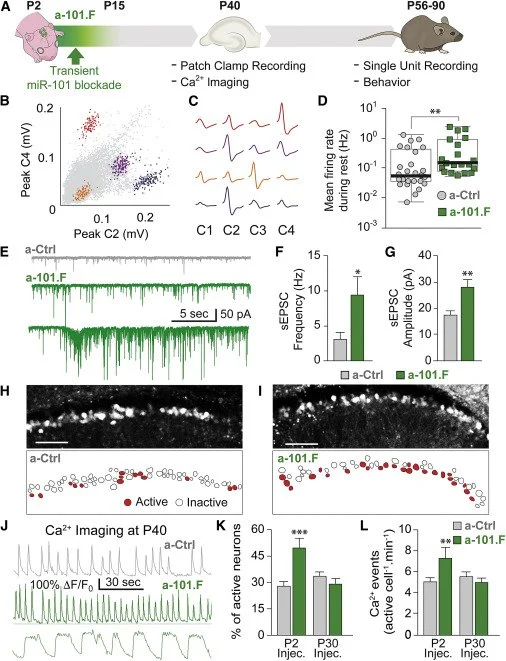
Our work has elucidated how microRNAs mediate various forms of neuronal plasticity, including changes in structure (Lippi, JCB 2011), synaptic strength (Saba, MCB 2012), and neurotransmitter identity (Dulcis*, Lippi*, Neuron 2017). We have discovered that microRNAs are master regulators of the complex series of developmental events that determine neural network stability in the adult (Lippi, Neuron 2016; Taylor, Elife 2023). We identified microRNA functions in vivo at multiple mechanistic levels, from molecular targets to neuronal structure, network activity in freely behaving animals, and behavior (Figure 3).
Figure 3. Transitory MiR-101 Inhibition Early in Development Induces Hyper-excitable Networks in the Young Adult. A. Schematic of the time course for miR-101 inhibition in vivo (green) and follow-up tests to assess consequences. B-C. Single-unit recordings from the hippocampus in freely behaving mice. D. Mean spike rates for single units in a-101.F- versus a-Ctrl-treated mice. E-G. sEPSC recordings in acute slices showing increased spontaneous excitatory activity in a-101.F-treated neurons. Spontaneous high-frequency burst discharges were seen in half of the slices from a-101.F-treated animals. H–L. Confocal imaging of spontaneous calcium transients on P40 showing increases in event frequency (center) and prolonged transients (bottom) in a-101.F-treated animals. From Lippi, Neuron 2016.
microRNAs Instruct Neuronal Diversification
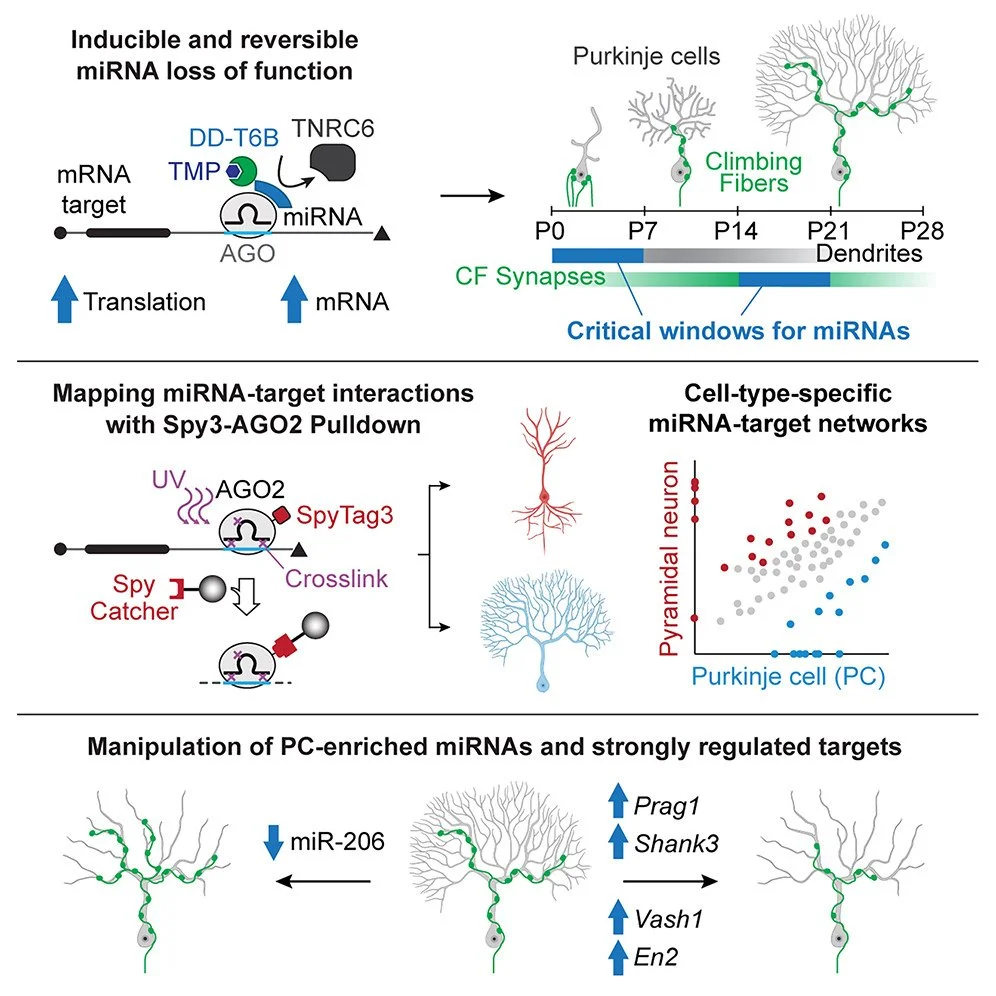
How did microRNAs help the emergence of complex brains? We hypothesize that they contributed to neuronal diversification. Purkinje Cells (PCs) are uniquely characterized by highly ramified, exuberant dendritic arbors. Are PC-specific microRNA programs driving this phenotype, which defines their identity? By comparing microRNA-target networks in developing PCs and pyramidal neurons (PNs), we identified microRNAs highly enriched in PCs but not in PNs and targets strongly repressed in PCs but not in PNs (despite the RNA being present in both neuronal subtypes). We found that miR-206, which is highly expressed in PCs, increases arborization in PNs (where it is normally absent) when overexpressed. Conversely, loss-of-function of miR-206 selectively in PCs strongly reduces their arborization. More importantly, we found that up-regulation in PCs of several targets that are specifically repressed in PCs but not in PNs (e.g., Prag1, Shank3, Vash1) strongly reduces PC dendritic arborization (Zolboot et al. Neuron 2025). Our findings indicate a prominent role for PC-specific microRNA-target networks in suppressing programs that restrict dendritogenesis. We demonstrated how differential repression of the same genes by microRNAs in separate neuronal subtypes leads to divergence in morphological features, suggesting that adding cell type-specific layers of post-transcriptional regulation on a common transcriptome could have been an evolutionary mechanism driving neuronal diversification.
Figure 4. Dissecting microRNAs roles critical for brain development. Top: We engineered technologies with vastly improved spatiotemporal resolution that allow the dissection of cell-type-specific miRNA-target networks. Fast and reversible miRNA loss of function showed that miRNAs are necessary for Purkinje cell (PC) differentiation, which previously appeared to be miRNA independent, and identified distinct critical miRNA windows for dendritogenesis and climbing fiber synaptogenesis, structural features defining PC identity. Middle: Using new mouse models that enable miRNA-target network mapping in rare cell types, we uncovered PC-specific post-transcriptional programs. Bottom: Manipulation of these programs revealed that the PC-enriched miR-206 and targets Shank3, Prag1, En2, and Vash1, which are uniquely repressed in PCs, are critical regulators of PC-specific dendritogenesis and synaptogenesis
For more information please check the PUBLICATIONS and NEWS pages.