Rooted in regulation: MicroRNAs as gardeners of the Purkinje cell dendritic arbor
/Cheryl Brandenburg∙ Roy V. Sillitoe
Abstract
Zolboot et al.1 developed tools to manipulate and map microRNA function in cerebellar Purkinje cells with temporal precision, revealing key roles in dendritic development and circuit connectivity. Their approach provides insight into microRNA-mediated regulation of neurodevelopment in sparse neuronal populations.
Is there any tree in our parks more elegant and leafier than the Purkinje corpuscle of the cerebellum … ? -Ramón y Cajal, 19172
Main text
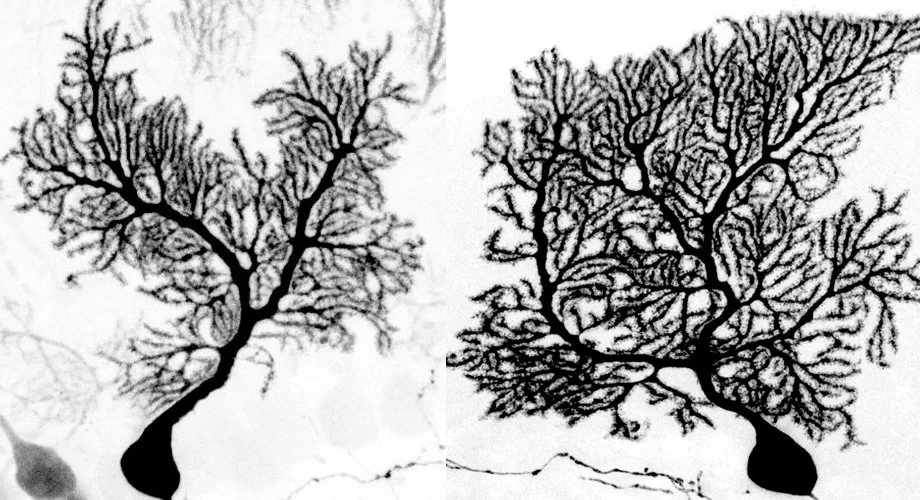
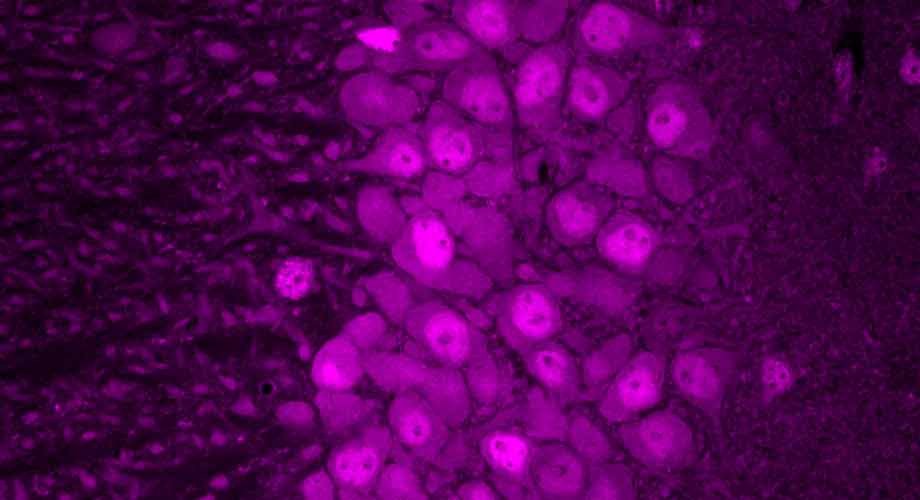
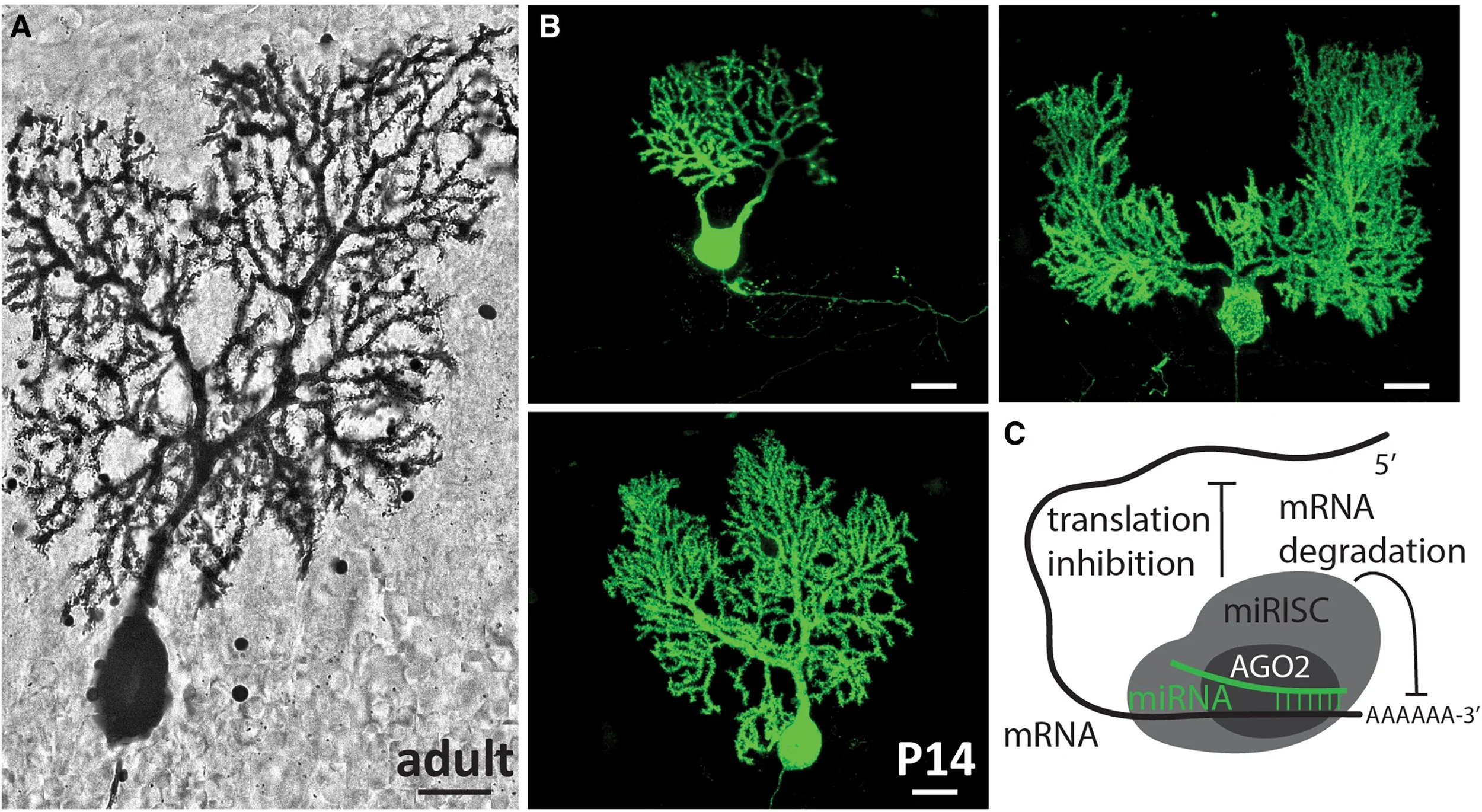
From the very first Golgi-stained images, the Purkinje cell dendritic arbor has awed neuroscientists with its striking morphology. Its expansive, planar branches spread like the limbs of an ancient tree, exquisitely sculpted to receive hundreds of thousands of synaptic inputs from parallel fibers, climbing fibers, and interneurons. Its sheer complexity and ordered architecture set it apart as a masterpiece of neural design. Just as a tree’s form is shaped by many forces—soil nutrients, seasonal cycles, fauna, and genetic blueprints—the Purkinje cell’s dendritic arbor emerges from an intricate molecular orchestration along a developmental timeline3 (Figure 1). Among the suspected architects of this process are microRNAs, small but powerful regulators that prune, refine, and guide neuronal structure. In this issue of Neuron, Zolboot et al.1 establish innovative techniques to uncover how microRNAs play a pivotal role in instructing Purkinje cell specification, shedding light on the molecular gardeners that cultivate one of the brain’s most elaborate neuronal landscapes.
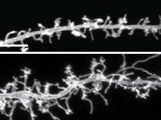
Figure 1 Purkinje cell dendrites are sculpted by a complex developmental program involving microRNAs
Purkinje cell development begins during mid-embryogenesis and continues over a particularly protracted time course through the first three postnatal weeks in the mouse.4 Tightly regulated gene expression programs establish cellular identity and connectivity within cerebellar microcircuits. Traditionally, neuronal subtype specification is attributed to transcription factor cascades that interact with local signaling cues. However, post-transcriptional regulatory mechanisms, such as those mediated by microRNAs, are emerging as important players in the precision and timing of these developmental programs. MicroRNAs are short (∼22 nucleotides), non-coding RNAs that repress target mRNAs through Argonaute (AGO) proteins (Figure 1 schematic), providing an additional layer of gene regulation that has expanded with evolution.5 Defining their precise roles in neuronal development—especially within specific cell types—has been limited by the lack of tools to manipulate microRNA activity with sufficient temporal precision. Most studies have relied on conditionally knocking out Dicer, which broadly and cumulatively disrupts canonical microRNA biogenesis over time. For example, targeted Dicer deletion in Purkinje cells leads to progressive cerebellar neurodegeneration and ataxia by around 13 weeks of age in mice.6 But could early developmental roles for microRNAs have gone undetected in such models, where microRNA loss causes defined defects only long after Dicer expression begins? To address this, Zolboot et al. developed a temporally restricted approach that preserves microRNA biogenesis while rapidly and reversibly inhibiting microRNA function using an updated version of the TNRC6B-derived peptide T6B. This strategy silences the mature microRNAs without altering their production, helping to avoid the confounding effects of redundancy or compensation that often accompany manipulation of individual microRNAs. As with a Dicer knockout approach, sustained loss of microRNA function led to Purkinje cell death, but with a markedly accelerated time course (by P28 rather than P90). Notably, when microRNA activity was transiently silenced early in brain development using the modified T6B peptide, Purkinje cells no longer underwent cell death. Instead, alterations in cell morphology, including changes in cell size and dendritic height, were observed. Combining different time points and technical modifications has therefore allowed for a more nuanced picture of the effect of microRNAs on the development of Purkinje cell morphology.
Another major challenge in microRNA research has been the identification of subtype-specific interactions, particularly in sparse populations, such as the Purkinje cells, which represent only ∼1% of cerebellar cells. To improve the mapping of microRNA-target interactions (MTIs) in rare, subtype-specific cell types, the authors developed an approach that circumvented limitations in available sequencing modalities through insertion of a small tag (SpyTag3) into an innocuous region of the endogenous Argonaute2 (Ago2) gene of the mouse. As a central component of microRNA function, here Ago2 serves to label mature associated microRNAs through SpyTag3, which the authors are able to then capture through cell-specific expression of SpyCatcher, enabling the resulting technique: Spy3-AGO2 pull-down and sequencing (SAPseq). By pulling down active microRNAs exclusively from Purkinje cells for sequencing and then comparing them to pyramidal neurons, they revealed that distinct sets of microRNAs act at specific stages of Purkinje cell development and were primarily related to dendritic growth, synapse formation, and neuronal maturation. They focused on two Purkinje cell-enriched microRNAs discovered by SAPseq and found that miR-206 manipulation reduced Purkinje cell complexity, much like their manipulations with the T6B peptide. Additionally, they uncovered that protein targets of the identified microRNAs were upregulated, namely, SHANK3, PRAG1, VASH1, and EN2. Importantly, several of these protein targets have well known roles during Purkinje cell development and cerebellar afferent fiber patterning with associated implications in neurological and neuropsychiatric conditions. Of particular interest, both Shank3 and En2 have been the focus in numerous mouse and human studies that link cerebellar deficits to autism spectrum disorders. If certain genes involved in autism lead to alterations in Purkinje cell dendritic complexity and disrupt cerebellar function, then altering Purkinje cell morphology directly should, at least to some extent, recapitulate distinct autism-associated behaviors. Indeed, after silencing microRNAs exclusively in Purkinje cells, here the authors uncovered deficits in motor coordination, sociability, hyperactivity, and stride length.
In addition to identifying several microRNAs that are responsible for dendritic growth, Zolboot et al. uncover how microRNAs shape climbing fiber connectivity with Purkinje cells. Climbing fibers provide powerful excitatory inputs to the cerebellum from the inferior olive. Climbing fiber innervation follows a strictly choreographed developmental timeline where multiple initial inputs are later pruned to a single dominant connection.7 SAPseq analysis revealed temporally dynamic microRNA expression patterns across phases of the climbing fiber pruning process that could be altered by perturbing microRNA function.
With this proof-of-concept approach for testing the role that microRNAs play in sculpting neuronal identity and morphology within a specific, sparse population, the techniques developed by Zolboot et al. provide a path for continued investigation of microRNA-mediated activity across other cell types, brain regions, and developmental contexts. Given the potential involvement of microRNA dysregulation in neurodevelopmental disorders such as autism8 and the ataxias9—both of which are linked to cerebellar function—methods that allow for more targeted dissection of microRNA function are essential for studying individual Purkinje cell subtypes in depth. Still underrepresented in the available strategies is a solution for better understanding specific Purkinje cell developmental programs. This hurdle exists due to the dynamic nature and heterogeneity of Purkinje cells within their functional “zonal” modules. Moreover, areas in the posterior cerebellum display significantly faster Purkinje cell maturational programs than Purkinje cells in the anterior cerebellum, which can be seen in the size of the dendritic tree.10 Therefore, Purkinje cells at the same developmental time point can diverge significantly from cells in a nearby area (Figure 1). Consequently, pull-down of microRNAs from all Purkinje cells may not discriminate between cells with very different rates of dendritic maturation. Clever use of the tools developed to express SpyCatcher within the highly heterogeneous subtypes of the Purkinje cell population could nevertheless enable more fine-tuned mapping of microRNA target interactions and how they contribute to the dramatic differences in dendritic development between cerebellar modules and lobules. These continued efforts may shed light on how specific genes are regulated between neuronal subpopulations in functional areas that can be targeted with treatments that are designed to differentially restore behaviors with precision.
In summary, SAPseq opens new avenues for exploring microRNA function in neuronal subtypes, illuminating post-transcriptional mechanisms that define cellular diversity in the nervous system. These small, yet potent, regulators influence Purkinje cell development at multiple levels, from promoting dendritic arborization to supporting synaptic elimination, refining our understanding of the intricate molecular choreography underlying the brain’s most elaborate dendritic architectures. Such insight will contribute to decoding how developmental programs optimize circuit function and shape the many animal behaviors.
Acknowledgments
This work was supported by Baylor College of Medicine, Texas Children’s Hospital, the National Institute of Neurological Disorders and Stroke (RVS: R01NS119301 and R01NS127435), and the Eunice Kennedy Shriver National Institute of Child Health and Human Development of the National Institutes of Health under Award Number P50HD103555 for use of the Cell and Tissue Pathogenesis Core (the BCM IDDRC).